torcheeg.model_selection¶
KFold¶
- class torcheeg.model_selection.KFold(n_splits: int = 5, shuffle: bool = False, random_state: int | None = None, split_path: None | str = None)[source][source]¶
A tool class for k-fold cross-validations, to divide the training set and the test set. One of the most commonly used data partitioning methods, where the data set is divided into k subsets, with one subset being retained as the test set and the remaining k-1 being used as training data. In most of the literature, K is chosen as 5 or 10 according to the size of the data set.
KFolddevides subsets without grouping. It means that during random sampling, adjacent signal samples may be assigned to the training set and the test set, respectively. When random sampling is not used, some subjects are not included in the training set. If you think these situations shouldn’t happen, consider usingKFoldPerSubjectGroupbyTrialorKFoldGroupbyTrial.
from torcheeg.model_selection import KFold from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFold(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
n_splits (int) – Number of folds. Must be at least 2. (default:
5)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
KFoldPerSubject¶
- class torcheeg.model_selection.KFoldPerSubject(n_splits: int = 5, shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool class for k-fold cross-validations, to divide the training set and the test set, commonly used to study model performance in the case of subject dependent experiments. Experiments were performed separately for each subject, where the data of the subject is divided into k subsets, with one subset being retained as the test set and the remaining k-1 being used as training data. In most of the literature, K is chosen as 5 or 10 according to the size of the data set.

from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.model_selection import KFoldPerSubject from torcheeg.utils import DataLoader cv = KFoldPerSubject(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.ToTensor(), transforms.To2d() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): # The total number of experiments is the number subjects multiplied by K train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
KFoldPerSubjectallows the user to specify the index of the subject of interest, when the user need to report the performance on each subject.from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.model_selection import KFoldPerSubject from torcheeg.utils import DataLoader cv = KFoldPerSubject(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset, subject=1): # k-fold cross-validation for subject 1 train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
n_splits (int) – Number of folds. Must be at least 2. (default:
5)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
KFoldCrossSubject¶
- class torcheeg.model_selection.KFoldCrossSubject(n_splits: int = 5, shuffle: bool = False, random_state: int | None = None, split_path: None | str = None)[source][source]¶
A tool class for k-fold cross-validations, to divide the training set and the test set. One of the most commonly used data partitioning methods, where the data set is divided into k subsets of subjects, with one subset subjects being retained as the test set and the remaining k-1 subset subjects being used as training data. In most of the literature, K is chosen as 5 or 10 according to the size of the data set.
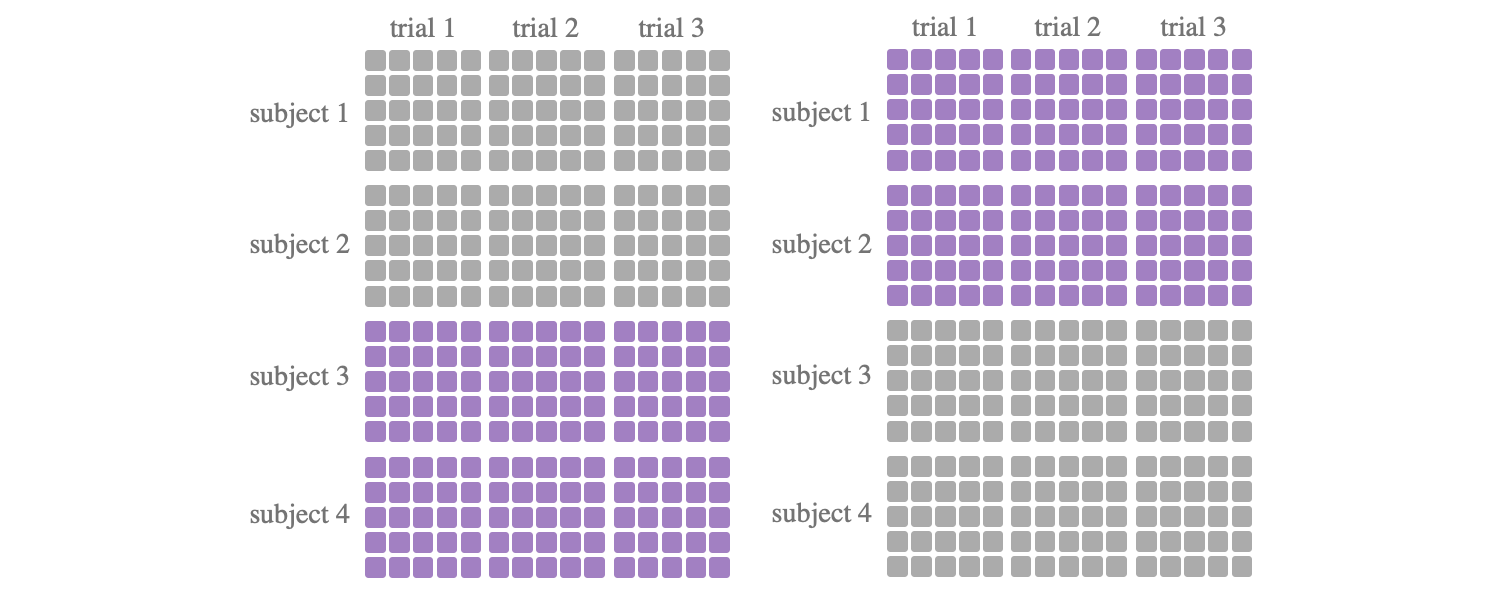
from torcheeg.model_selection import KFoldCrossSubject from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFoldCrossSubject(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
n_splits (int) – Number of folds. Must be at least 2. (default:
5)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
KFoldGroupbyTrial¶
- class torcheeg.model_selection.KFoldGroupbyTrial(n_splits: int = 5, shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool class for k-fold cross-validations, to divide the training set and the test set. A variant of
KFold, where the data set is divided into k subsets at the dimension of trials, with one subset being retained as the test set and the remaining k-1 being used as training data. In most of the literature, K is chosen as 5 or 10 according to the size of the data set.KFoldGroupbyTrialdevides subsets at the dimension of trials. Take the first partition withk=5as an example, the first 80% of samples of each trial are used for training, and the last 20% of samples are used for testing. It is more consistent with real applications and can test the generalization of the model to a certain extent.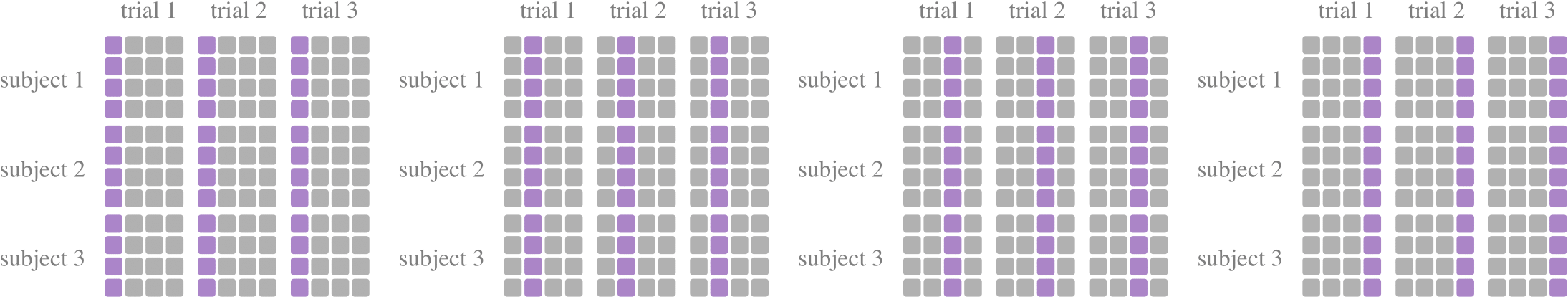
from torcheeg.model_selection import KFoldGroupbyTrial from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFoldGroupbyTrial(n_splits=5, shuffle=False) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
n_splits (int) – Number of folds. Must be at least 2. (default:
5)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
KFoldPerSubjectGroupbyTrial¶
- class torcheeg.model_selection.KFoldPerSubjectGroupbyTrial(n_splits: int = 5, shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool class for k-fold cross-validations, to divide the training set and the test set, commonly used to study model performance in the case of subject dependent experiments. Experiments were performed separately for each subject, where the data for all trials of the subject is divided into k subsets at the trial dimension, with one subset being retained as the test set and the remaining k-1 being used as training data. In most of the literature, K is chosen as 5 or 10 according to the size of the data set.

from torcheeg.model_selection import KFoldPerSubjectGroupbyTrial from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFoldPerSubjectGroupbyTrial(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.ToTensor(), transforms.To2d() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): # The total number of experiments is the number subjects multiplied by K train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
KFoldPerSubjectGroupbyTrialallows the user to specify the index of the subject of interest, when the user need to report the performance on each subject.from torcheeg.model_selection import KFoldPerSubjectGroupbyTrial from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFoldPerSubjectGroupbyTrial(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset, subject=1): # k-fold cross-validation for subject 1 train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
n_splits (int) – Number of folds. Must be at least 2. (default:
5)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
KFoldCrossTrial¶
- class torcheeg.model_selection.KFoldCrossTrial(n_splits: int = 5, shuffle: bool = False, random_state: int | None = None, split_path: None | str = None)[source][source]¶
A tool class for k-fold cross-validations, to divide the training set and the test set. One of the most commonly used data partitioning methods, where the data set is divided into k subsets of trials, with one subset trials being retained as the test set and the remaining k-1 subset trials being used as training data. In most of the literature, K is chosen as 5 or 10 according to the size of the data set.
KFoldCrossTrialdevides subsets at the dataset dimension. It means that during random sampling, adjacent signal samples may be assigned to the training set and the test set, respectively. When random sampling is not used, some subjects are not included in the training set. If you think these situations shouldn’t happen, consider usingKFoldPerSubjectGroupbyTrialorKFoldGroupbyTrial.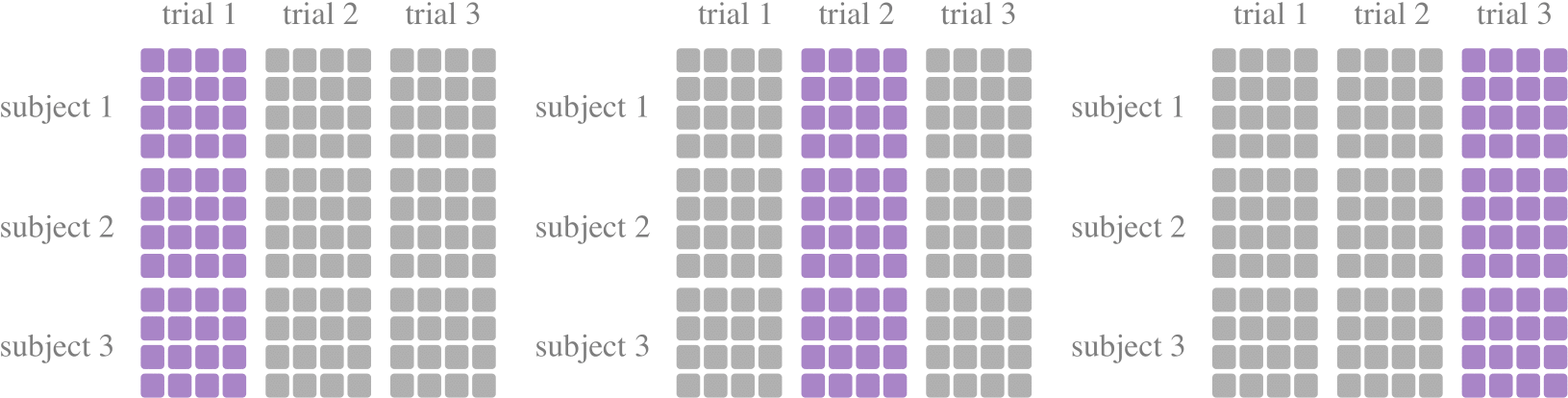
from torcheeg.model_selection import KFoldCrossTrial from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFoldCrossTrial(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
n_splits (int) – Number of folds. Must be at least 2. (default:
5)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
KFoldPerSubjectCrossTrial¶
- class torcheeg.model_selection.KFoldPerSubjectCrossTrial(n_splits: int = 5, shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool class for k-fold cross-validations, to divide the training set and the test set, commonly used to study model performance in the case of subject dependent experiments. Experiments were performed separately for each subject, where the data set is divided into k subsets of trials, with one subset trials being retained as the test set and the remaining k-1 subset trials being used as training data. In most of the literature, K is chosen as 5 or 10 according to the size of the data set.
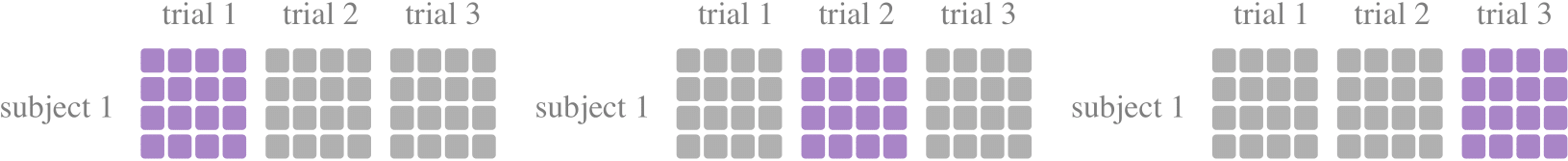

from torcheeg.model_selection import KFoldPerSubjectCrossTrial from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFoldPerSubjectCrossTrial(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.ToTensor(), transforms.To2d() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): # The total number of experiments is the number subjects multiplied by K train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
KFoldPerSubjectCrossTrialallows the user to specify the index of the subject of interest, when the user need to report the performance on each subject.from torcheeg.model_selection import KFoldPerSubjectCrossTrial from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = KFoldPerSubjectCrossTrial(n_splits=5, shuffle=True) dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset, subject=1): # k-fold cross-validation for subject 1 train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
n_splits (int) – Number of folds. Must be at least 2. (default:
5)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
LeaveOneSubjectOut¶
- class torcheeg.model_selection.LeaveOneSubjectOut(split_path: None | str = None)[source][source]¶
A tool class for leave-one-subject-out cross-validations, to divide the training set and the test set, commonly used to study model performance in the case of subject independent experiments. During each fold, experiments require testing on one subject and training on the other subjects.
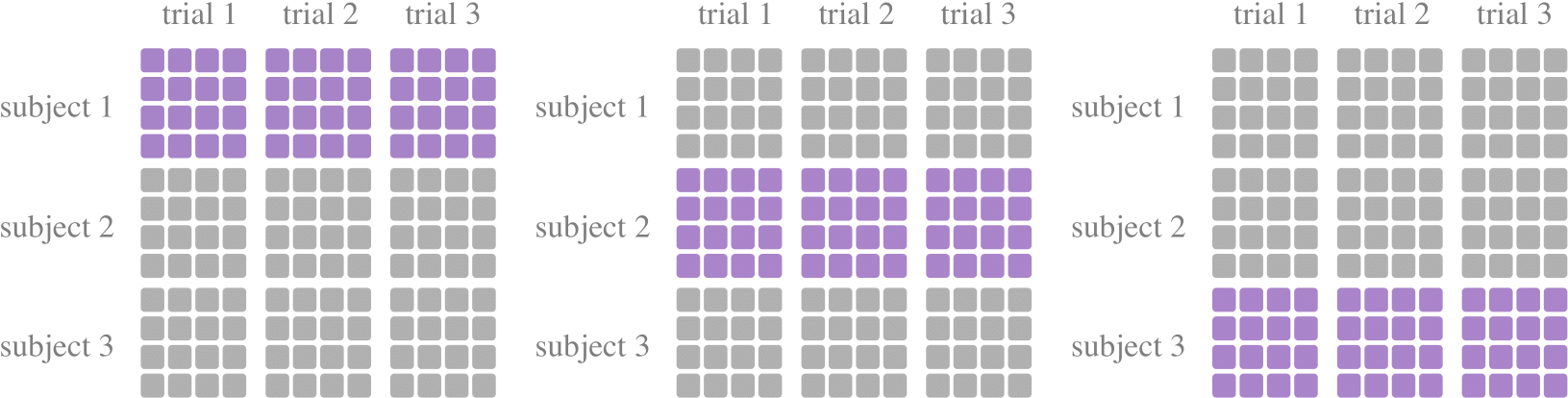
from torcheeg.model_selection import LeaveOneSubjectOut from torcheeg.datasets import DEAPDataset from torcheeg import transforms from torcheeg.utils import DataLoader cv = LeaveOneSubjectOut() dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.ToTensor(), transforms.To2d() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) for train_dataset, test_dataset in cv.split(dataset): train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
Subcategory¶
- class torcheeg.model_selection.Subcategory(criteria: str = 'task', split_path: None | str = None)[source][source]¶
A tool class for separating out subsets of specified categories, often used to extract data for a certain type of paradigm, or for a certain type of task. Each subset in the formed subset list contains only one type of data.
Common usage:
from torcheeg.datasets import M3CVDataset from torcheeg.model_selection import Subcategory from torcheeg import transforms from torcheeg.utils import DataLoader cv = Subcategory() dataset = M3CVDataset(root_path='./aistudio', online_transform=transforms.Compose( [transforms.To2d(), transforms.ToTensor()]), label_transform=transforms.Compose([ transforms.Select('subject_id'), transforms.StringToInt() ])) for subdataset in cv.split(dataset): loader = DataLoader(subdataset) ...
TorchEEG supports the division of training and test sets within each subset after dividing the data into subsets. The sample code is as follows:
cv = Subcategory() dataset = M3CVDataset(root_path='./aistudio', online_transform=transforms.Compose( [transforms.To2d(), transforms.ToTensor()]), label_transform=transforms.Compose([ transforms.Select('subject_id'), transforms.StringToInt() ])) for i, subdataset in enumerate(cv.split(dataset)): train_dataset, test_dataset = train_test_split(dataset=subdataset, split_path=f'./split{i}') train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
For the already divided training and testing sets, TorchEEG recommends using two
Subcategoryto extract their subcategories respectively. On this basis, thezipfunction can be used to combine the subsets. It is worth noting that it is necessary to ensure that the training and test sets have the same number and variety of classes.train_cv = Subcategory() train_dataset = M3CVDataset(root_path='./aistudio', online_transform=transforms.Compose( [transforms.To2d(), transforms.ToTensor()]), label_transform=transforms.Compose([ transforms.Select('subject_id'), transforms.StringToInt() ])) val_cv = Subcategory() val_dataset = M3CVDataset(root_path='./aistudio', subset='Calibration', num_channel=65, online_transform=transforms.Compose( [transforms.To2d(), transforms.ToTensor()]), label_transform=transforms.Compose([ transforms.Select('subject_id'), transforms.StringToInt() ])) for train_dataset, val_dataset in zip(train_cv.split(train_dataset), val_cv.split(val_dataset)): train_loader = DataLoader(train_dataset) val_loader = DataLoader(val_dataset) ...
- Parameters:
criteria (str) – The classification criteria according to which we extract subsets of data for the including categories. (default:
'task')split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
train_test_split¶
- torcheeg.model_selection.train_test_split(dataset: BaseDataset, test_size: float = 0.2, shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool function for cross-validations, to divide the training set and the test set. It is suitable for experiments with large dataset volume and no need to use k-fold cross-validations. The test samples are sampled according to a certain proportion, and other samples are used as training samples. In most literatures, 20% of the data are sampled for testing.
train_test_splitdevides the training set and the test set without grouping. It means that during random sampling, adjacent signal samples may be assigned to the training set and the test set, respectively. When random sampling is not used, some subjects are not included in the training set. If you think these situations shouldn’t happen, consider usingtrain_test_split_per_subject_groupby_trialortrain_test_split_groupby_trial.
from torcheeg.datasets import DEAPDataset from torcheeg.model_selection import train_test_split from torcheeg import transforms from torcheeg.utils import DataLoader dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) train_dataset, test_dataset = train_test_split(dataset=dataset) train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
dataset (BaseDataset) – Dataset to be divided.
test_size (int) – If float, should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the test split. If int, represents the absolute number of test samples. (default:
0.2)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
train_test_split_groupby_trial¶
- torcheeg.model_selection.train_test_split_groupby_trial(dataset: BaseDataset, test_size: float = 0.2, shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool function for cross-validations, to divide the training set and the test set. It is suitable for experiments with large dataset volume and no need to use k-fold cross-validations. The test samples are sampled according to a certain proportion, and other samples are used as training samples. In most literatures, 20% of the data are sampled for testing.
train_test_split_groupby_trialdevides training set and the test set at the dimension of each trial. For example, whentest_size=0.2, the first 80% of samples of each trial are used for training, and the last 20% of samples are used for testing. It is more consistent with real applications and can test the generalization of the model to a certain extent.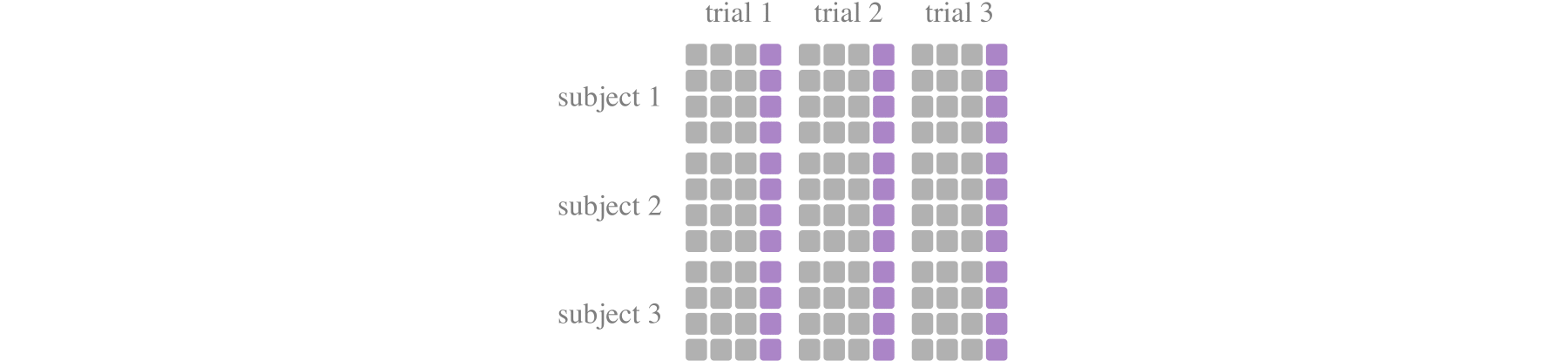
from torcheeg.datasets import DEAPDataset from torcheeg.model_selection import train_test_split_groupby_trial from torcheeg import transforms from torcheeg.utils import DataLoader dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) train_dataset, test_dataset = train_test_split_groupby_trial(dataset=dataset) train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
dataset (BaseDataset) – Dataset to be divided.
test_size (int) – If float, should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the test split. If int, represents the absolute number of test samples. (default:
0.2)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
train_test_split_per_subject_groupby_trial¶
- torcheeg.model_selection.train_test_split_per_subject_groupby_trial(dataset: BaseDataset, test_size: float = 0.2, subject: str = 's01.dat', shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool function for cross-validations, to divide the training set and the test set. It is suitable for subject dependent experiments with large dataset volume and no need to use k-fold cross-validations. For the first step, the EEG signal samples of the specified user are selected. Then, the test samples are sampled according to a certain proportion for each trial for this subject, and other samples are used as training samples. In most literatures, 20% of the data are sampled for testing.

from torcheeg.datasets import DEAPDataset from torcheeg.model_selection import train_test_split_per_subject_groupby_trial from torcheeg import transforms from torcheeg.utils import DataLoader dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.ToTensor(), transforms.To2d() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) train_dataset, test_dataset = train_test_split_per_subject_groupby_trial(dataset=dataset) train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
dataset (BaseDataset) – Dataset to be divided.
test_size (int) – If float, should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the test split. If int, represents the absolute number of test samples. (default:
0.2)subject (str) – The subject whose EEG samples will be used for training and test. (default:
s01.dat)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
train_test_split_cross_trial¶
- torcheeg.model_selection.train_test_split_cross_trial(dataset: BaseDataset, test_size: float = 0.2, shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool function for cross-validations, to divide the training set and the test set. It is suitable for experiments with large dataset volume and no need to use k-fold cross-validations. Parts of trials are sampled according to a certain proportion as the test dataset, and samples from other trials are used as training samples. In most literatures, 20% of the data are sampled for testing.
train_test_split_cross_trialdevides training set and the test set at the dimension of each trial. For example, whentest_size=0.2, the first 80% of samples of each trial are used for training, and the last 20% of samples are used for testing. It is more consistent with real applications and can test the generalization of the model to a certain extent.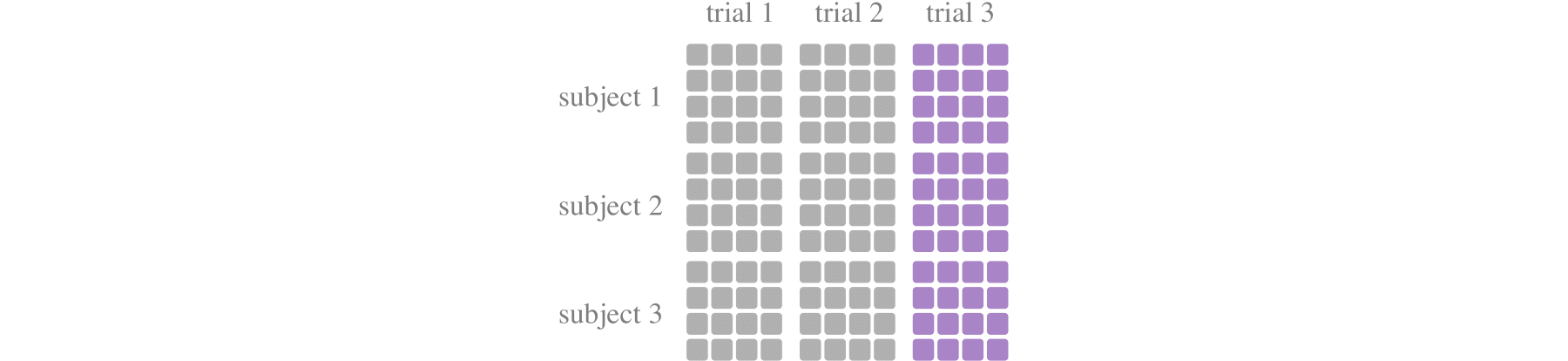
from torcheeg.datasets import DEAPDataset from torcheeg.model_selection import train_test_split_cross_trial from torcheeg import transforms from torcheeg.utils import DataLoader dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.To2d(), transforms.ToTensor() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) train_dataset, test_dataset = train_test_split_cross_trial(dataset=dataset) train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
dataset (BaseDataset) – Dataset to be divided.
test_size (int) – If float, should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the test split. If int, represents the absolute number of test samples. (default:
0.2)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
train_test_split_per_subject_cross_trial¶
- torcheeg.model_selection.train_test_split_per_subject_cross_trial(dataset: BaseDataset, test_size: float = 0.2, subject: str = 's01.dat', shuffle: bool = False, random_state: float | None = None, split_path: None | str = None)[source][source]¶
A tool function for cross-validations, to divide the training set and the test set. It is suitable for subject dependent experiments with large dataset volume and no need to use k-fold cross-validations. For the first step, the EEG signal samples of the specified user are selected. Then, parts of trials are sampled according to a certain proportion as the test dataset, and samples from other trials are used as training samples. In most literatures, 20% of the data are sampled for testing.

from torcheeg.datasets import DEAPDataset from torcheeg.model_selection import train_test_split_per_subject_cross_trial from torcheeg import transforms from torcheeg.utils import DataLoader dataset = DEAPDataset(root_path='./data_preprocessed_python', online_transform=transforms.Compose([ transforms.ToTensor(), transforms.To2d() ]), label_transform=transforms.Compose([ transforms.Select(['valence', 'arousal']), transforms.Binary(5.0), transforms.BinariesToCategory() ])) train_dataset, test_dataset = train_test_split_per_subject_cross_trial(dataset=dataset) train_loader = DataLoader(train_dataset) test_loader = DataLoader(test_dataset) ...
- Parameters:
dataset (BaseDataset) – Dataset to be divided.
test_size (int) – If float, should be between 0.0 and 1.0 and represent the proportion of the dataset to include in the test split. If int, represents the absolute number of test samples. (default:
0.2)subject (str) – The subject whose EEG samples will be used for training and test. (default:
s01.dat)shuffle (bool) – Whether to shuffle the data before splitting into batches. Note that the samples within each split will not be shuffled. (default:
False)random_state (int, optional) – When shuffle is
True,random_stateaffects the ordering of the indices, which controls the randomness of each fold. Otherwise, this parameter has no effect. (default:None)split_path (str) – The path to data partition information. If the path exists, read the existing partition from the path. If the path does not exist, the current division method will be saved for next use. If set to None, a random path will be generated. (default:
None)
